|
Problem 60: Minimal Absent Words
|

|
Words that do not occur in files provide a useful technique to discard or
discriminate files.
They act like the set of factors of a file but admit a
more compact trie representation as factor codes.
A word $w$, $|w|\gt 1$, is said to be absent or forbidden in a word $x$ if it
does not occur in $x$.
It is said to be minimal with this property if in
addition both $w[0\dd |w|-2]$ and $w[1\dd |w|-1]$ do occur in $x$.
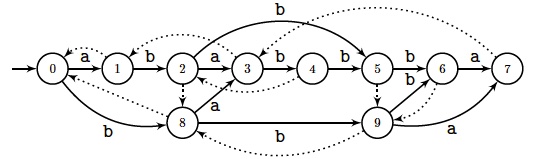
A natural method to compute the minimal words absent in $x$ is to start with
its Factor automaton $\aFact(x)$, smallest deterministic automaton accepting
all its factors.
Each factor is spelt out along a path starting from the
initial state and all states are terminal.
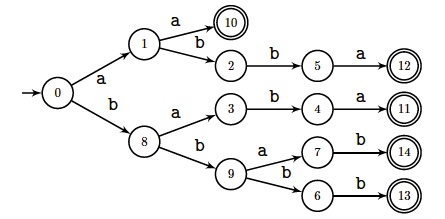
Design a linear-time algorithm to compute the trie of minimal absent words of
a word $x$ from its Factor automaton $\aFact(x)$.
Use failure links of the automaton.
Design a linear-time algorithm to recover the Factor automaton of a word $x$
from the trie of its minimal absent words.
Use the Aho-Corasick technique.
References
A. V. Aho and M. J. Corasick. Efficient string matching: An aid to
bibliographic search. Commun. ACM, 18(6):333-340, 1975.
M. Béal, F. Mignosi, and A. Restivo. Minimal forbidden words and
symbolic dynamics. In C. Puech and R. Reischuk, editors, STACS 96,
13th Annual Symposium on Theoretical Aspects of Computer Science,
Grenoble, France, February 22-24, 1996, Proceedings, volume 1046 of
Lecture Notes in Computer Science, pages 555-566. Springer, 1996.
M.-P. Béal, M. Crochemore, F. Mignosi, A. Restivo, and M. Sciortino.
Forbidden words of regular languages. Fundam. Inform., 56(1,2):121-
135, 2003.
A. Blumer, J. Blumer, D. Haussler, A. Ehrenfeucht, M. T. Chen, and
J. I. Seiferas. The smallest automaton recognizing the subwords of a
text. Theor. Comput. Sci., 40:31-55, 1985.
S. Chairungsee and M. Crochemore. Using minimal absent words to
build phylogeny. Theor. Comput. Sci., 450(1):109-116, 2012.
M. Crochemore. Transducers and repetitions. Theor. Comput. Sci.,
45(1):63-86, 1986.
M. Crochemore, C. Hancart, and T. Lecroq. Algorithms on Strings.
Cambridge University Press, 2007. 392 pages.
M. Crochemore, A. Héliou, G. Kucherov, L. Mouchard, S. P. Pissis, and
Y. Ramusat. Absent words in a sliding window with applications. Inf.
Comput., 270, 2020.
M. Crochemore, F. Mignosi, and A. Restivo. Automata and forbidden
words. Inf. Process. Lett., 67(3):111-117, 1998.
M. Crochemore, F. Mignosi, A. Restivo, and S. Salemi. Text compression
using antidictonaries. In J. Wiedermann, P. van Emde Boas, and
M. Nielsen, editors, International Conference on Automata, Languages
an Programming (Prague, 1999), number 1644 in LNCS, pages 261-270.
Springer-Verlag, 1999. Rapport I.G.M. 98-10, Université de Marne-la-
Vallée.
M. Crochemore and W. Rytter. Text algorithms. Oxford University
Press, 1994. 412 pages.
M. Crochemore and W. Rytter. Jewels of Stringology. World Scientific
Publishing, Hong-Kong, 2002. 310 pages.
T. Mieno, Y. Kuhara, T. Akagi, Y. Fujishige, Y. Nakashima, S. Inenaga,
H. Bannai, and M. Takeda. Minimal unique substrings and minimal
absent words in a sliding window. In A. Chatzigeorgiou, R. Dondi,
H. Herodotou, C. A. Kapoutsis, Y. Manolopoulos, G. A. Papadopoulos,
and F. Sikora, editors, SOFSEM 2020: Theory and Practice of Computer
Science - 46th International Conference on Current Trends in Theory
and Practice of Informatics, SOFSEM 2020, Limassol, Cyprus, January
20-24, 2020, Proceedings, volume 12011 of Lecture Notes in Computer
Science, pages 148-160. Springer, 2020.
T. Ota, H. Fukae, and H. Morita. Dynamic construction of an antidictionary
with linear complexity. Theor. Comput. Sci., 526:108-119,
2014.
T. Ota and H. Morita. On a universal antidictionary coding for stationary
ergodic sources with finite alphabet. In International Symposium
on Information Theory and its Applications, ISITA 2014, Melbourne,
Australia, October 26-29, 2014, pages 294-298. IEEE, 2014.
T. Ota and H. Morita. A compact tree representation of an antidictionary.
IEICE Transactions, 100-A(9):1973-1984, 2017.
R. M. Silva, D. Pratas, L. Castro, A. J. Pinho, and P. J. S. G. Ferreira.
Three minimal sequences found in ebola virus genomes and absent from
human DNA. Bioinformatics, 31(15):2421-2425, 2015.